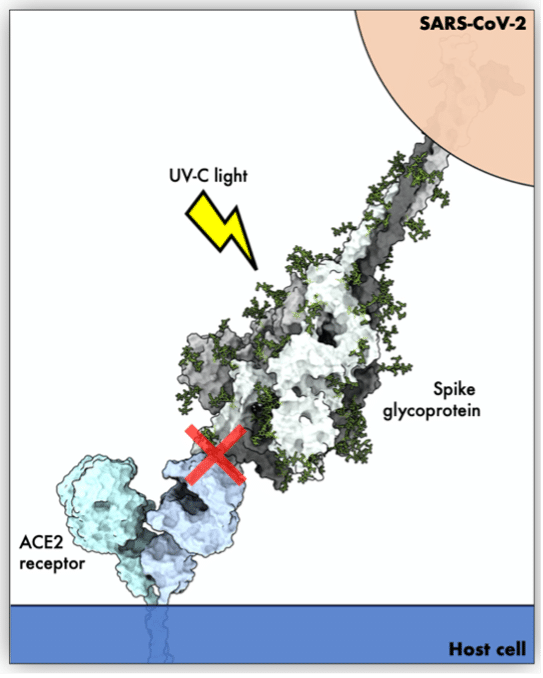
This success story deals with antiviral and virucidal activities against SARS-CoV-2 and how researchers can improve the development of antiviral agents by gaining insight into the molecular mechanism of spike protein inactivation after UV-C exposure.
About
The National Research Council (CNR) is the largest public research institution in Italy performing multidisciplinary research activities.
Founded in 1923, CNR’s mission is to perform research in its own Institutes, to promote innovation and competitiveness of the national industrial system, to promote the internationalisation of the national research system, to provide technologies and solutions to emerging public and private needs, to advise Government and other public bodies, and to contribute to the qualification of human resources.

In 2021, a consortium of three Italian research teams working at the Institute of Biomedical Technologies of the National Research Council (CNR-ITB), at the University of Brescia (UniBs), and the University of Genoa (UniGe) received support from EGI to support investigational studies on fully-glycosylated full-length SARS-CoV-2 spike protein.
In a nutshell, the main objective of this research activity was to characterise the molecular mechanism of spike protein inactivation and identification of antiviral scaffolds promising against SARS-CoV2. What the three Italian research teams do specifically is to:
- Calculate dynamic molecular trajectories resulting from simulations of the use case that will be shared with the scientific community through public repositories (data delivery).
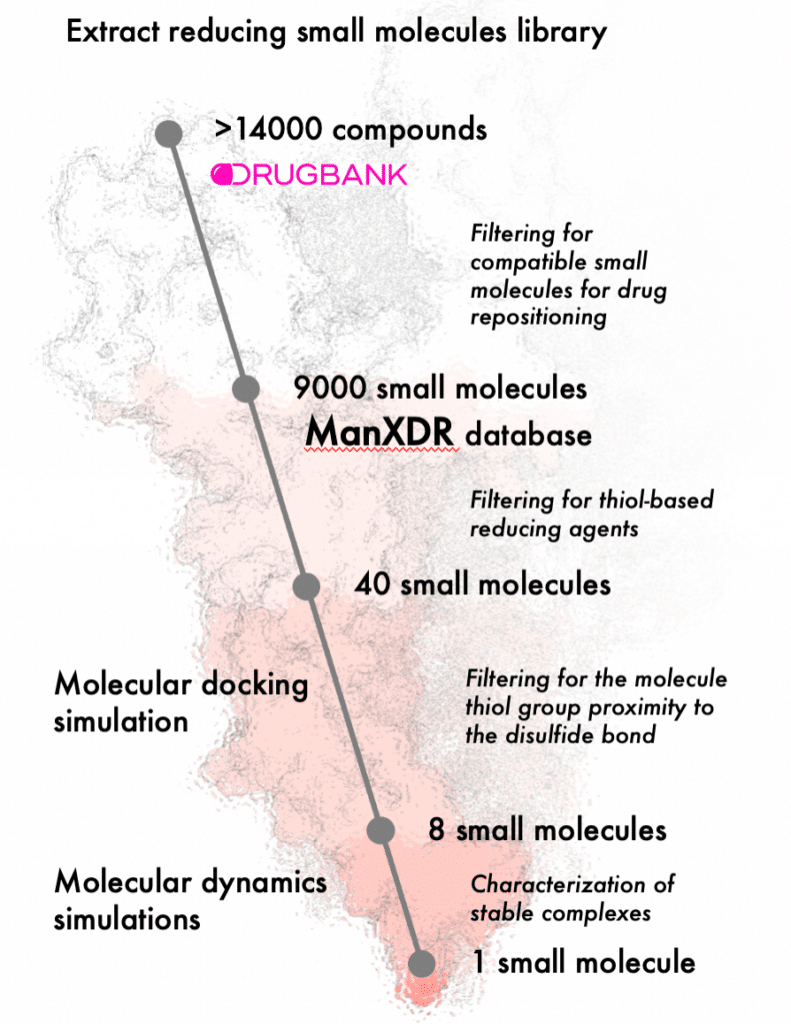
- Run drug repositioning pipeline for identification of reducing agents against SARS-CoV2.
- Implement AI and HPC-based solutions to analyse biomedical data.
- Organise advanced training courses in drug discovery studies and in advanced bio simulations for young scientists: M.S. Thesis and PhD students.
The insights gained from this computational study could provide key atomistic details to design spike protein inactivators to develop new agents against SARS-CoV-2.
The challenge
The Researchers’ team has already performed molecular dynamics studies to evaluate S1 subunit variants, they are located in the receptor binding domain (RBD) and near the insertion 680SPRRA R↓SV687 forming a cleavage motif RxxR for furin-like enzymes. Comparative studies between wt and variants highlighted some critical regions for repositioning known drugs already available on the market.
Although the community already used GPU devices in previous molecular dynamics studies, they proved to be inadequate to deal with the use case at hand efficiently. The main limitations were related to their computation power and their memory size, which did not permit them to fit the entire computation on the GPU memory, consistently reducing the performance.
Modern GPU cards made available by EGI have overcome these limitations by significantly optimising the performance and, consequently, time-to-solution. The technical support and the resource pool offered by EGI played an active role in guaranteeing exhaustive research of the conformational space of the spike protein. This exploration is mandatory to define the effect of UV-C light on the Spike protein, and thus it helps in the identification of novel antiviral drugs specifically targeting SARS-CoV2.
The implemented use case was aimed at assessing whether the molecular mechanisms of spike protein inactivation after UV-C radiation can assist drug discovery in the identification of specific anti-SARS-CoV2 drugs.
To this end, 16 molecular dynamics simulations were performed on compound-spike protein complexes selected in a previous analysis performed by the research teams for 11 microseconds.
Performing these dynamic simulations is very challenging as it requires very powerful computational resources (i.e., GPU devices exploiting the most recent architectures) and a lot of disk space to store the results of the simulations.
EGI provided ‘free at the point of use’ computing resources to implement the analysis. Moreover, the shepherd supported us in the implementation of the use case and in addressing specific computational challenges.
The solution
The team used the following EGI services:
- The EGI Cloud Compute and EGI Online Storage to distribute the computational task to a scalable compute platform.
- The EGI Check-In to enable user registration, authenticated and authorised access to the underlying distributed compute infrastructure.
- Technical consultancy and assistance to profit from the EGI solutions.
Services provided by EGI
Impact
registered in the vo.i-nergy.eu
consumed over the last two years
consumed over the last two years
consumed over the last two years
used to perform molecular dynamics simulations
Supporting projects
EGI-ACE is a 30-month project with a mission to empower researchers from all disciplines to...